How EU projects work on supercomputing applications to help contain the corona virus pandemic - MaX contribution
The Centres of Excellence in high-performance computing are working to improve supercomputing applications in many different areas: from life sciences and medicine to materials design, from weather and climate research to global system science. A hot topic that affects many of the above-mentioned areas is, of course, the fight against the corona virus pandemic.
There are rather obvious challenges for those EU projects that are developing HPC applications for simulations in medicine or in the life sciences, like CompBioMed (Biomedicine) BioExcel (Biomolecular Research), and PerMedCoE (Personalized Medicine). But also other projects from different scientific areas, that you would, at first sight, not directly relate to research on the pandemic, are developing and using appropriate applications to model the virus and its spread, and support policy makers with computing-heavy simulations.
MaX CoE is one of these. Our engagement is in Modelling the electronic structure of the protease.
Though the main interest of the MaX flagship codes is then centered on materials science, the CoE is participating in the fight against SARS-CoV-2. Given the critical pandemic situation that the world is currently facing, an unprecedented effort is being devoted to the study of SARS-CoV-2 by researchers from different scientific communities and groups worldwide. From the biomolecular standpoint, particular focus is being devoted to the main protease, as well as to the spike protein. As such, it is an important potential antiviral drug target: if its function is inhibited, the virus remains immature and non-infectious. Using fragment-based screening, researchers have identified a number of small compounds that bind to the active site of the protease and can be used as a starting point for the development of protease inhibitors.
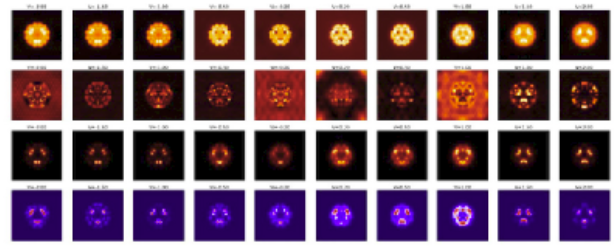
Sars-Cov-2 main protease monomer, in green, with the N3 3-mer peptide inhibitor bound in the enzyme’s active site.(from PDB crystal structure 6lu7). Structure like this ones can be simulated with a full DFT calculation and automatically decomposed into fragments whose interaction network can be characterized and analyzed.
Among other quantities, MaX researchers now have the possibility to model the electronic structure of the protease in contact with a potential docked inhibitor, and provide new insights on the interactions between them by selecting specific amino-acids that are involved in the interaction and characterizing their polarities. This new approach proposed by the MaX scientists is complementary to the docking methods used up to now and based on in-silico research of the inhibitor. Biological systems are naturally composed of fragments such as amino-acids in proteins or nitrogenous bases in DNA.
With this approach, it is possible to evaluate whether the amino acid-based fragmentation is consistent with the electronic structure resulting from the QM computation. This is an important indicator for the end-user, as it enables to evaluate the quality of the information associated with a given fragment. Then, QM observables on the system’s fragments can be obtained, which are based on a population analysis of electronic density of the system, projected on the amino-acid.
A novelty that this approach enables is the possibility of quantifying the strength of the chemical interaction between the different fragments. It is possible to select a target region and identify which fragments of the systems interact with this region by sharing electrons with it.
“We can reconstruct the fragmentation of the system in such a way as to focus on an active site in a specific portion of the protein”, says Luigi Genovese from CEA (Commissariat à l’énergie atomique et aux énergies alternatives) who is heading Max’s efforts on this topic. “We think this modelling approach could inform efforts in protein design by granting access to variables otherwise impervious to observation.”
Additional Information:
- Microscopic Factors Modulating the Interactions Between the SARS-CoV-2 Main Protease and α−Ketoamide Inhibitors, Genovese et al., preprint (2020)
- Polaris (MD)
- BigDFT
- MaX Webinar: “The Flexibilities of Wavelets for Electronic Structure Calculations in Large Systems”
To have a complet overview over the various ways in which EU projects are using supercomputing applications to tackle and support the global challenge of containing the pandemic, please visit FocusCoE news section.